- Published:
- 16 April 2026
- Author:
- Max Jackson and Professor Sarah Coupland
- Read time:
- 10 Mins
Max Jackson and Professor Sarah Coupland report on the use of AI in detecting uveal melanoma and metastatic carcinomas to the choroid. AI was applied to both clinical and cytopathological images, aiming to enable earlier and more accurate diagnosis of these challenging ocular lesions. Max Jackson joined the Liverpool Ocular Oncology Research Group (LOORG) as a PhD student to carry out this work.
Uveal melanoma (UM) is the most common primary intraocular malignancy in adults, but it is nonetheless very rare, affecting approximately 6 people per million annually.1 Despite its rarity, UM is deadly; it spreads to the liver in around 50% of cases, with most patients surviving <6 months post-metastasis.2,3
Detection is challenging because UM shares clinical features with other ocular lesions. Choroidal freckles (naevi) are small, pigmented lesions in the choroid that rarely cause harm. While approximately 1 in 9,000 naevi transform into UM,4 most remain stable and require only routine monitoring by an optician. Conversely, metastatic carcinomas of the choroid (mCA), which primarily spread from breast and lung cancers to the eye, are the most common intraocular malignancy.
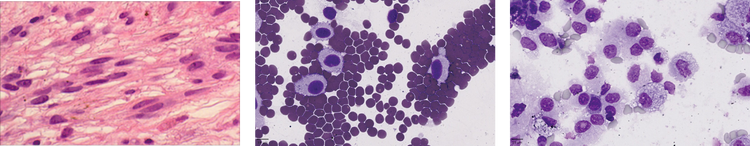
Both naevi and mCA create diagnostic challenges in primary care and specialist ocular centres alike. At the Liverpool Ocular Oncology Centre (LOOC), patients may undergo intraocular biopsy, in difficult cases where imaging alone cannot establish an unequivocal diagnosis (Figures 1a–c). Cytospins of the biopsies are prepared and reviewed by our specialist pathologist for pathological confirmation (Figures 1e and f).1
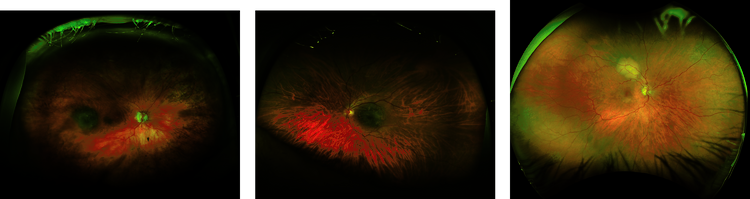
Most naevi are initially detected by opticians, who refer patients to local eye hospitals or specialist centres for further assessment. A combination of 3 non-invasive imaging modalities (fundoscopy, optical coherence tomography and ultrasound) are used to characterise a lesion. Figures 1a–c show fundoscopy images for naevi, UM and mCA. In most cases, this allows a relatively straightforward diagnosis that does not require the patient to be seen by a specialist ocular oncologist. Naevi constitute a significant proportion of referrals to LOOC, increasing waiting times for UM and mCA patients where timely diagnosis is critical.
For malignant lesions seen at the LOOC, UM and mCA differentiation can pose a challenge to even specialist clinicians: most UM are pigmented but many are not, and virtually all mCA are also non-pigmented. If there is insufficient diagnostic information from the imaging modalities alone, biopsies can improve diagnostic accuracy. The biopsy procedure itself is highly specialised and performed at only a handful of centres across the globe, while pathologists with expertise in diagnosing these rare ocular cancers are equally scarce. Therefore, there is a challenge in differentiating, both using non-invasive images and cytology. Very occasionally, a choroidal naevus can be an incidental finding in an enucleated globe (Figure 1d).
In summary, the 3 challenges when it comes to diagnosis of ocular lesions are: naevus or UM; UM or mCA on imaging modalities; and UM or mCA on cytology whole slide images (WSI). All of these issues have the potential to be addressed with AI.
Starting with the non-invasive fundoscopy images, 6 AI architectures (RETFound,5 ResNet50,6 EfficientNet-b0,7 DenseNet121,8 MobileNetv29 and VGG1610) were applied to differentiate first between naevi and UM and then between UM and mCA. These architectures were selected for their different advantages, with some being less computationally intensive and others better at pattern detection. We trained these models on images from 2,456 UM patients, 1,810 naevus patients and 66 mCA patients.
Cytology WSIs require a different processing approach, due to their high resolution and large file sizes. We first divided each WSI into thousands of small tiles, automatically removing those with minimal tissue content (>95% white space).
We then trained a U-Net, which is a type of AI model designed specifically for segmentation tasks, to identify and outline individual tumour cells. Using a small subset of manually annotated tiles containing both UM and mCA cells, this segmentation model achieved 99% accuracy in detecting cancer cells. Once trained, we applied it across all patient tiles, retaining only those containing 5 or more segmented tumour cells. These tumour-containing tiles were then classified by 5 of our AI architectures (RETFound is a fundoscopy-specific architecture) to differentiate between UM and mCA cells.
Classification performance was measured using area under the curve (AUC) of the receiver operator characteristic curve (ROC). An AUC of 0.50 is equivalent to a random guess, with anything >0.90 showing excellent performance and 1.00 being the perfect classifier.
The models demonstrated excellent diagnostic performance across all 3 classification tasks. For naevi versus UM differentiation, RETFound achieved an AUC of 0.90, crossing the desired threshold. The UM versus mCA classification achieved even better performance, with RETFound, DenseNet121 and MobileNetv2 all achieving a near-perfect performance of 0.99.
Finally, the most challenging task is differentiating between UM- and mCA- tumour cells on the cytology images. VGG16 achieved an AUC of 0.96, again demonstrating its excellent discriminatory power.
While these results show very high accuracy, there are some limitations that need to be addressed. Obtaining data in rare cancers is difficult, due to the limited number of centres seeing these patients and to the ethical restrictions on accessing data from other centres. We have begun to address this issue with our new trusted research environment, EYE-CAN-AID Bioresource, which contains clinical data, ocular images, radiological images, histopathological WSIs and genetic data. However, external validation from a separate centre would be a required as a next step in testing these models.
As with all AI models, there’s the challenge of the ‘black box’ problem, where we have no true understanding of why models make particular decisions. One approach to address this is via class activation maps (CAM), which generate heat maps highlighting the image regions that are the most influential in the model’s classification (decision making). In our case, Grad-CAM11 was applied to the cytospin WSIs. Figure 2 shows an example of a segmented cytospin tile by our model alongside a CAM for the same tiles. The model focused on specific tumour cells (shown in red/yellow) while correctly classifying this tile as a UM. However, it is not immediately clear why some cells were more important than others in this decision.
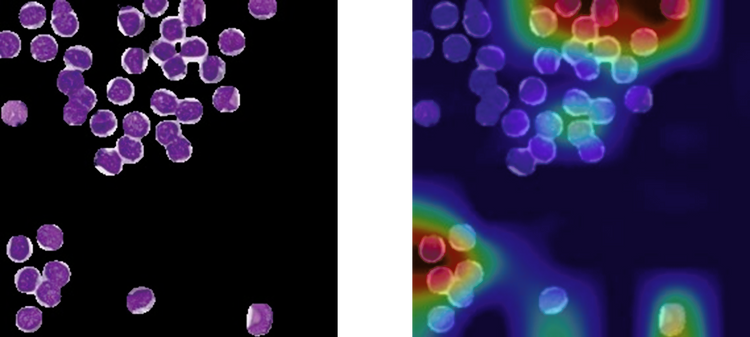
Figure 2: Example of the same tile of a UM patient, pre- and post-classification. (a) Original tile post U-Net segmentation. (b) CAM tile after classification. Red is the region most used by the architecture.
There are many instances of studies similar to ours that boast great performance and make a great research paper.12 However, this is often where these projects end, with no real-world change to help the clinician or patient. Why is this? Firstly, there are obvious regulatory considerations that need to be addressed. The process of getting an AI model into routine use is very unclear, with no clear set guidance in the UK, making it difficult to even know where to start this process. Additionally, the point at which a model is deemed good enough to be implemented is also not defined – that is, at what point is the model considered okay and how many tests should be performed first?
Another key consideration is integration into existing workflows. Are these models quick and easy to use? Do they have clear instructions for both technical and non-technical staff? Is there infrastructure in place to host AI models? When performing these studies, typically a high-end computer and advanced coding knowledge is required; these models are not finished products for use by anyone.
Finally, do clinicians want or need this? Establishing ground truth for successful diagnosis is challenging in rare disease centres where specialist expertise is limited. Comparing performance with and without AI support becomes nearly impossible when the clinicians themselves represent the gold standard.
Collaborations between AI researchers and clinicians are needed to identify real-world problems that can be solved with the help of AI and then fully adopted into routine clinical pathways. Researchers must not only focus on achieving high accuracy, but also on practical implementations developed alongside frequent consultations with the end users.
Clinicians, meanwhile, should remain open to new approaches that could improve patient diagnosis and prognosis, even if it possibly means changing established workflows. Digitisation of pathology is no small undertaking, but it is essential if we are to test these AI models in real-world practice and ultimately improve outcomes for cancer patients, including those with rare tumours.
Acknowledgments
The authors would like to thank the whole team of the Liverpool Ocular Oncology Centre, and Dr Helen Kalirai, Custodian of the Liverpool Ocular Oncology Bioresource (including EYE-CAN-AID).
References available on our website.