The current expansion in genomic testing aims to realise the long-promised potential of personalised medicine. There is hope that this will impact on cancer survival statistics when compared to similarly developed nations across Europe and beyond, in addition to promoting equity, access and efficiency.1 This genomic revolution, spearheaded in the UK by the 100,000 Genomes Project and currently being actuated by the NHS Genomic Medicine Service (GMS),2 will effectively drive the use of genomic testing within the NHS for years to come.
It has been argued that, of all the laboratory-based specialties, pathology has been slow to embrace the emergence of the genomic era.3 However, in many respects, pathologists have been performing genomic analysis using surrogate markers for decades, so this integration of knowledge is not new. While a major component of the GMS revolves around investigation of so-called rare disease, it is in the field of cancer that the impact on pathology will be most encountered.
Given the progress of translational research in yielding real-world results, we are now rapidly approaching a crossroads – genomic medicine will change pathology, but how will that change manifest in the coming years? And critically, how will pathology handle the multiple challenges of expanding genomic testing – best use of tissue, ensuring high-quality samples for testing, interpretation and integration of results, knowledge expansion, and workforce and capacity issues – to deliver on the GMS and its potential?
Guardians of the tissue
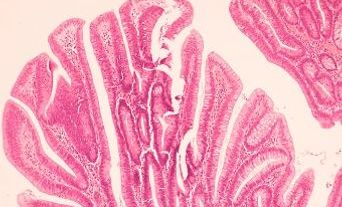
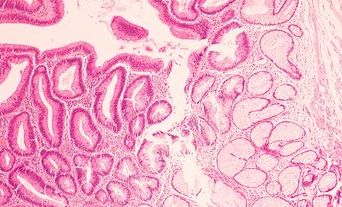
As the first port of call for cells and tissue taken during investigations for suspected cancers, pathologists have long been the gateway to testing, determining the best use of this limited resource. It seems reasonable to assume that next generation sequencing (NGS)-based testing will require pathological input, although the extent of involvement will depend upon the specialty proactively seeking a pivotal role.
The pathologist is key in assessing the sample and tumour content and predicting which samples are likely to be insufficient for panel testing.
In a drive to improve turnaround times of genomic tests, it has been suggested that clinicians could send tissue directly for genomic testing, for example colorectal biopsies, for mismatch repair testing. Furthermore, some tests are not dependent on the final histopathological diagnosis. One such recent development is NICE guidance on NTRK testing and treatment, now approved for use in specific circumstances but histology-agnostic,4 so testing could proceed in parallel with histological assessment. Any tumour positive for the fusion protein will be eligible for treatment with entrectinib and larotrectinib.5 However, as any pathologist will recognise, there is no guarantee that any given biopsy will contain cancer cells or an adequate ratio of cancer to normal tissue, both of which would have significant implications for test result interpretation.
Further challenges also include ensuring that, where tissue is particularly limited, at least the minimum required genomic testing is achieved and not sacrificed at the expense of a failed panel test. Here again, the pathologist is key in assessing the sample and tumour content and predicting which samples are likely to be insufficient for panel testing. Local systems should be in place for more limited testing – a so-called ‘salvage pathway’ – so patient care is not compromised. Thus, management of the testing process is most effectively undertaken by the pathologist. In keeping with this, some Genomics Laboratory Hubs’ websites clearly state that the pathologist will determine the test of choice and results will be returned to them.6
A fixation on formalin
Clearly, morphological analysis remains the core of cancer diagnosis, with additional tests being done largely to confirm and refine diagnosis and provide prognostic and/or predictive information. Thus, formalin fixation of tissues to achieve optimal morphological quality has been the mainstay of histopathology. However, genomic testing is more accurate and requires lower input when fresh frozen (FF) samples are used rather than formalin-fixed paraffin embedded tissue. Indeed, whole genome sequencing (WGS) via the 100,000 Genomes Project showed improved single nucleotide variation and copy number alteration calling when performed on FF tissue, as described by Arumugan et al.7 Within the GMS, FF tissue for WGS is mandatory.
Formaldehyde is now defined as a Carc1B (presumed to have carcinogenic potential for humans),8 and recent EU law reduced allowable workplace exposure from 2 ppm to 0.5 ppm.9,10 This has encouraged attempts to move away from formalin in theatres and reduce exposure within the laboratory by trialling systems of vacuum-packed specimen transfer or use of alternatives fixatives. These two elements combined are a powerful incentive to reduce formalin use despite the very obvious upfront costs and requirement for new specimen pathways.
Adoption of alternative fixatives is challenging in terms of cost, safety, efficacy and compatibility with established immunohistochemical (IHC) and in situ hybridisation protocols.11 Paxgene is one such option and has been shown to provide better DNA and RNA integrity for molecular testing, but provides poorer H&E and IHC staining for higher cost.12 Recently, evidence suggested that the reduction in concentration of neutral buffered formalin (NBF), when compared with higher concentrations of NBF or 10% formal saline for routine fixation, may increase DNA yield without impacting morphology.13
It seems unlikely we will see a dramatic shift away from formalin in the near term, although with the expansion of WGS in the GMS there will be pressure for laboratories to receive tissue fresh with the onus on labs to process, sample for genomic testing and fix safely, which may drive a need for out-of-hours working patterns. Increased handling of fresh tissue may bring with it risk of infection and it seems unlikely that formalin will be totally removed from the process owing to its usefulness as an antimicrobial,14 but parallel systems to achieve the goals of the GMS may need to be developed.
NGS technologies
There is an ever-increasing array of technologies and approaches to genomic testing, as well as the bioinformatics and data science tools used to analyse outputs. While it is unlikely that pathologists will be responsible for primary interpretation of genomic tests, knowledge of the technologies applied will be important to understand their limitations in the context of an integrated pathology report.
Most genomic testing focuses on sequencing either DNA or RNA, or methylation analysis, and are built on the work of Frederick Sanger’s sequencing by synthesis.15 This technique is still commercially available and gives accurate reads of 600–1,000 bases in length.16 However, it is not high throughput, which led to the development of multiple NGS techniques in an effort to drive down price and increase speed. Here, DNA or RNA is extracted, fragmented, prepared with specific adaptors (‘library prep’) and then sequenced. This results in millions of smaller reads rather than fewer, longer reads, but at the cost of accuracy.
There are many different techniques currently employed for sequencing, with the choice of platform largely dictated by desired experimental outcomes.
Errors are mitigated by the large number of reads generated by individual fragments of DNA combined with statistical correction. Some areas remain difficult to sequence owing to homopolymeric regions (consecutive identical bases), GC bias,17 and centromeric or telomeric regions.16
There are many different techniques currently employed for sequencing, with the choice of platform largely dictated by desired experimental outcomes. These include Ion Torrent, Illumina, Pacific Biosciences (PacBio) and Oxford Nanopore.18–21 Ion Torrent measures hydrogen ions released when a complementary nucleotide is incorporated into a growing DNA chain.22
By contrast, Illumina binds DNA fragments to a glass slide and sequential addition of proprietary nucleotides with fluorescent reporters occurs, with the reported colour corresponding to the base added.23 The process is repeated up to 300-times, either from one end of a DNA fragment (single-read) or both ends (paired-end). Both Ion Torrent and Illumina result in relatively short lengths of DNA read in each run, but many millions of fragments of DNA can be read simultaneously.
PacBio and Oxford Nanopore are both longread sequencers. PacBio can read fragments up to 30–50 kb in length, using an engineered DNA polymerase bound to the DNA and anchoring it within a microscopic well. Nucleotides with fluorophores are incorporated and recorded in real time owing to the unique properties of the circular DNA template and well construction on the flow cell.21 Oxford Nanopore uses a membrane with protein pores that bind a polymerase and ratchet through DNA as nucleotides are added. Detection of the current change across the membrane enables base calling.24 This system allows ultralong reads (longest to date >4 Mb) with a generally higher error rate than short reads, but with the ability to sequence long, repetitive regions.
Currently, the GMS uses the Illumina platform and Genomics England Bioinformatics pipeline for commissioned WGS (at present including paediatric tumours, sarcomas and some haematological malignancies) with large NGS cancer panel testing recommended for all other solid tumours. The knowledge gap posed by this new technology is well recognised by NHS England, Health Education England and the College, and a range of approaches are being taken to address this, as outlined by Mahamdallie et al.25 While this education will be embedded in the curriculum of our trainees, a more bespoke approach will need to be taken to ensure the upskilling of our existing consultant workforce.
Shaping the future of pathology
The desire for economies of scale, standardisation and equity of access to genomic testing within the NHS has led to the centralisation of purported genomic testing within England at seven NHS Genomic Laboratory Hubs.26 These hubs will be linked to the National Genomic Test Directory,27 listing the full range of tests funded by the NHS in England. In relation to cancer, these are divided into solid tumours (adult), neurological tumours, sarcomas, haematological tumours and paediatric tumours. It is expected there will be equity of access for all those eligible to get a test, with standardisation across the country.
The College’s view is that ‘pathologists are at the heart of these developments because of their vast experience in tissue handling, processing and reporting’,28 which accords with many of the key challenges involved in the successful implementation of the GMS. The government published Genome UK: The Future of Healthcare in September last year,29 and set out ‘the vision to extend the UK’s leadership in genomic healthcare and research’. Part of that vision is to link genomic data with medical records and digital pathology information, with the realisation that much work will be required ‘to meet the needs of the pathology workforce to support cancer genomics’.
The College’s view is that ‘pathologists are at the heart of these developments because of their vast experience in tissue handling, processing and reporting’ ... which accords with many of the key challenges involved in the successful implementation of the GMS.
The aim of the NHS Genomic Medicine Service is to ‘offer genomic testing routinely to all people with cancer’,2 so it is clear that sampling, tumour assessment, sequencing and analysis of cancers across the NHS will be routine in the near future. There is little doubt this will drive advances in molecular classification and subtyping of cancers given the vast amount of data that global sequencing within a national healthcare service will provide when linked to clinical data and long-term outcomes. It will also provide prognostic and therapeutic data relating to specific genomic changes such as drug targetability or response to immunomodulation.
Routine genomic analysis on this scale is unprecedented and will place the UK at the forefront of precision cancer diagnostics. But pathology is key to the realisation of this vision, not only in ensuring delivery of testing, but also in the collation of this data with morphological features, which will offer a unified pathological answer to the clinical question.